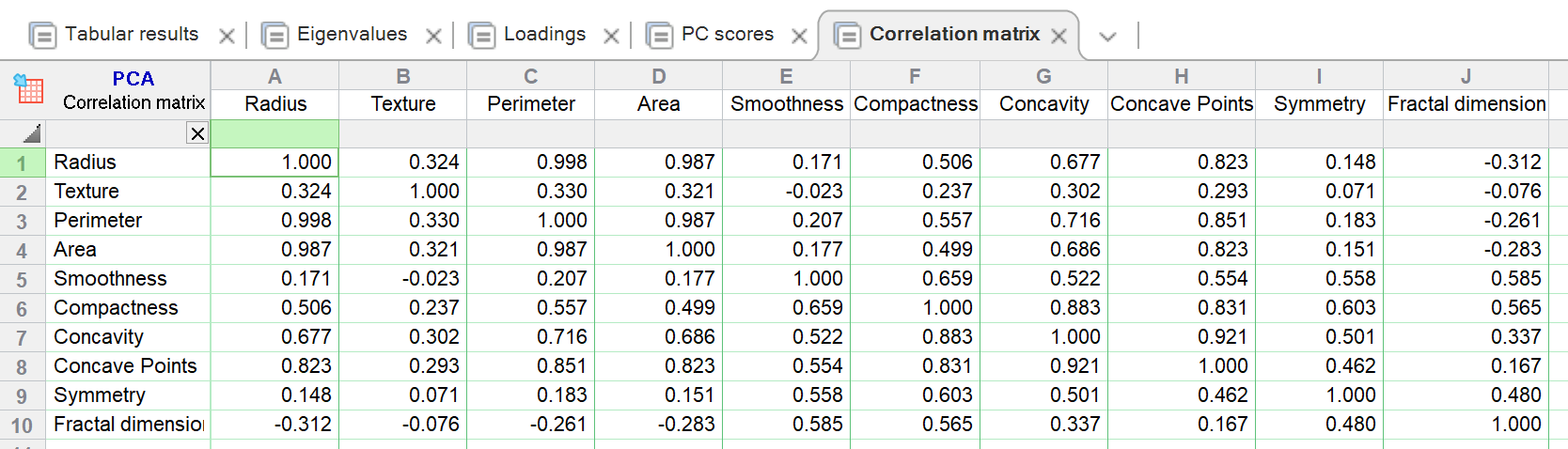
This scaling is also known as maximum likelihood scaling (MALS). In MLPCA one applies a scaling (or even a rotation) such that the measurement errors in the measured variables are independent and distributed according to the standard normal distribution. I personally find it very valuable to discuss these options in light of the maximum-likelihood principal component analysis model (MLPCA). Hence, my answer is to use covariance matrix when variance of the original variable is important, and use correlation when it is not. 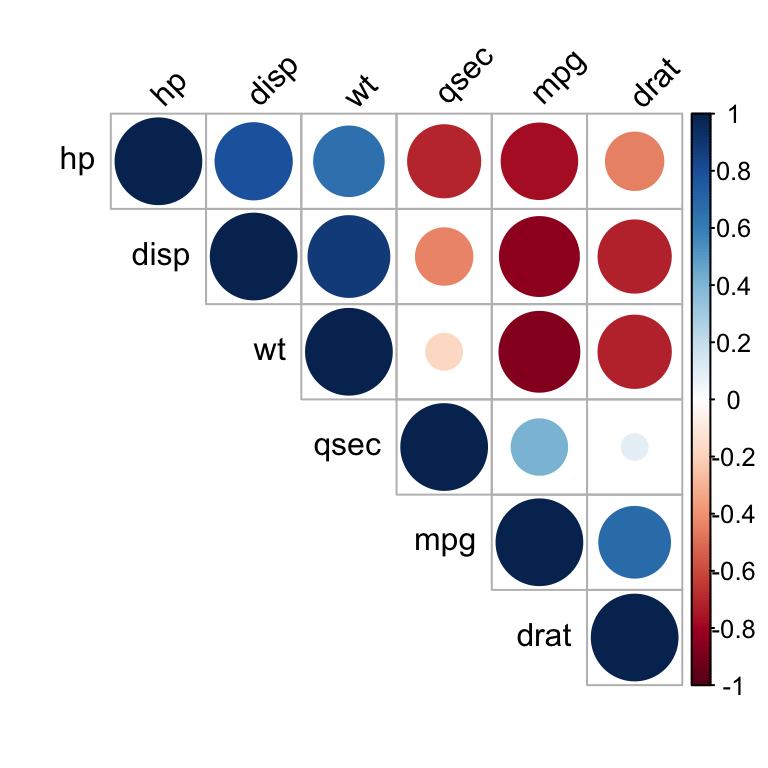
Using the covariance matrix is one way for building factors that account for the size of the state. In some applications we do want to adjust for the size of the state. Should we use correlation matrix here? It depends. Now, the scales could be different: DC has 600K and CA - 38M people. The units of measure are the same - counts (number) of people. It shouldn't matter whether you use meters or centimeters in this case, so you could argue that correlation matrix should be used.Ĭonsider now population of people in different states. Imagine that your variables have different units of measure, such as meters and kilograms. Otherwise, why would anyone ever do covariance PCA? It would be safer to always perform correlation PCA. However, this is only true when scale of the variables isn't a factor. The advantage of using VDW scores is that skewness and outlier effects are removed from the data, and can be used if the goal is to perform an analysis under the contraints of normality - and every variable needs to be purely standard normal distributed with no skewness or outliers.Ī common answer is to suggest that covariance is used when variables are on the same scale, and correlation when their scales are different. Use of VDW scores is very popular in genetics, where many variables are transformed into VDW scores, and then input into analyses. You actually don't need to think about the difference of using the correlation matrix $\mathbf(0.975)$. gene expression from the same platform with similar range and scale, or you are working with log equity asset returns, then using correlation will throw out a tremendous amount of information. However, if all of your data are based on e.g. UNTRANSFORMED (RAW) DATA: If you have variables with widely varying scales for raw, untransformed data, that is, caloric intake per day, gene expression, ELISA/Luminex in units of ug/dl, ng/dl, based on several orders of magnitude of protein expression, then use correlation as an input to PCA. Notice also that the outlying individuals (in this data set) are outliers regardless of whether the covariance or correlation matrix is used. PCA on correlation is much more informative and reveals some structure in the data and relationships between variables (but note that the explained variances drop to $64\%$ and $71\%$). Notice that PCA on covariance is dominated by run800m and javelin: PC1 is almost equal to run800m (and explains $82\%$ of the variance) and PC2 is almost equal to javelin (together they explain $97\%$). 
Hep.PC.cov = prcomp(heptathlon, scale=FALSE) Hep.PC.cor = prcomp(heptathlon, scale=TRUE) Now let's do PCA on covariance and on correlation: # scale=T bases the PCA on the correlation matrix This outputs: hurdles highjump shot run200m longjump javelin run800m Heptathlon # look at heptathlon data (excluding 'score' variable) Some of the variables have an average value of about 1.8 (the high jump), whereas other variables (run 800m) are around 120. Especially when the scales are different.Īs an example, take a look at this R heptathlon data set. In general, PCA with and without standardizing will give different results. Using the correlation matrix is equivalent to standardizing each of the variables (to mean 0 and standard deviation 1). You tend to use the covariance matrix when the variable scales are similar and the correlation matrix when variables are on different scales.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
